Abstract
This study investigated the human risk of infection due to inadvertent ingestion of water during swimming in a river that receives SARS-CoV-2-containing effluent from a wastewater treatment plant (WWTP). A quantitative microbial risk assessment (QMRA) approach was applied for risk estimation using dose-response models (DRM) of different surrogate coronaviruses (SARS-CoV-1, MERS-CoV) and the virus responsible for most infectious respiratory illnesses (i.e., influenza A H5N1) due to the unavailability of DRM for SARS-CoV-2. The ratio of infectious concentration to genomic copies of SARS-CoV-2 is unknown and also unavailable for other coronaviruses. Therefore, literature-based information on enteric viruses was used for formulating the ratio used for QMRA, although it is acknowledged that identifying this information for SARS-CoV-2 is a priority, and in the absence of information specific to SARS-CoV-2, another coronavirus would be a preferable surrogate to the enteric viruses used here. The calculated concentration of ingested SARS-CoV-2 ranged between 4.6 × 10−7 and 80.5 genomic copies/dip (one swim = 32 mL). The risk of infection (> 9 × 10−12 to 5.8 × 10−1) was found to be > 1/10,000 annual risk of infection. Moreover, the study revealed that the risk estimation was largely dependent on the value of the molecular concentration of SARS-CoV-2 (gc/mL). Overall immediate attention is required for obtaining information on the (i) ratio of infectious virus to genomic copies, (ii) DRM for SARS-CoV-2, and (iii) virus reduction rate after treatment in the WWTPs. The QMRA structure used in present findings is helpful in analyzing and prioritizing upcoming health risks due to swimming performed in contaminated rivers during the COVID-19 outbreak.
Graphical abstract

Similar content being viewed by others
Introduction
China reported an outbreak of SARS-CoV-2 (disease known as COVID-19) in late December 2019 of unidentified etiology occurring in Wuhan, Hubie Province, to the World Health Organization (World Health Organization 2020a). The typical presentation of this disease includes dry cough, fatigue, fever, and myalgia, and more than half of the patients developed dyspnea (Chen et al. 2020; Guan et al. 2020; Huang et al. 2020; Wang et al. 2020a, b). WHO declared it is a global pandemic and a public health emergency of international concern on January 31, 2020 (World Health Organization 2020b). It is believed that the major transmission routes are inhalation via person-to-person and aerosol/droplet transmission and/or personal contact with contaminated surfaces, hand-facilitated transfer of the virus from contaminated objects to the mouth, nose, or eyes (Haas 2020). SARS-CoV-2 and its surrogates can survive on different surfaces and water matrices from days to months (Table S1 in supporting information) although its infectivity potential and persistence have not been fully understood. The SARS-CoV-2 and its viral RNA are shed from bodily excreta, such as urine and feces of COVID-19-positive patients and subsequently find a way to wastewater treatment plants (WWTPs, Table 1) (Ahmed et al. 2020; Kitajima et al. 2020; Nemudryi et al. 2020; Randazzo et al. 2020; Rimoldi et al. 2020; Wu et al. 2020). Medema et al. (2020) reported the first study of detection of SARS-CoV-2 in raw sewage samples from seven different cities and the Netherlands, airport. They noted that SARS-CoV-2 was not detected in the samples collected 3 weeks before the first COVID-19 case; meanwhile, the first RNA virus was detected in sewage at five sites (WWTPs), exactly 1 week after the first COVID-19 case. Thereafter, Wu et al. (2020) measured the virus from the urban treatment facility located in Massachusetts (USA) and noted 2 × 102 genomic copies/mL in raw wastewater. From Australia (Queensland), Ahmed et al. (2020) noted the value of virus RNA in raw wastewater that ranged between 0.019 and 1.2 genomic copies/mL. Further, a study conducted by Xiao et al. (2020) reported the infectious virus in the urine sample and feces of COVID-19-positive patients and concluded that fecal-oral transmission could be an additional route for virus spread. However, detailed literature is not available about the infectious concentration of SARS-CoV-2 in the saliva, urine, and feces or raw wastewater. Without knowing the infectious concentration of the virus, it is difficult to understand the pathogenicity and risk of infection. Researchers reported that 1000 genomic copies of Adenovirus 40/41 are equivalent to 01 PFU (infectious virus) (He and Jiang 2005; Aslan et al. 2011; McBride et al. 2013; Carducci et al. 2016). However, information is missing about the coronaviruses in this regard and genomic copies to PFU correlation was largely dependent on the viral strains. Moreover, a dose-response model (DRM) is currently unavailable for estimating the risk of infection due to SARS-CoV-2 (Table S2 in supporting material indicates the available DRM for SAR-COV-1, SARS, and its surrogate viruses). Wu et al. (2020) noted that 41 out of 74 COVID-19 patients had SARS-CoV-2 RNA-positive fecal samples for a mean of 27.9 days, whereas the respiratory samples of these patients remained positive for a mean of 16.7 days. One patient had RNA-positive fecal samples for 33 days after their respiratory samples became RNA-negative, and another patient tested RNA-positive in their fecal sample for 47 days after symptom onset. Similar results were cited by Ng and Tilg (2020) and concluded that extended shedding of the virus in the fecal samples of COVID-19 patients was observed in spite of the absence of the virus in the respiratory tract and nasal swab samples. They suggested stool testing of hospitalized patients even after recovery in order to avoid the community spread (fecal-oral transference).
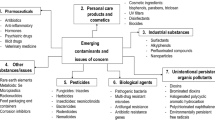
A waterborne transmission has never been reported in literature; however, their high abundance in feces and urine may contribute (Ahmed et al. 2020; Heller et al. 2020; Kitajima et al. 2020). Also, past history shows that faulty sewage pipelines (aerosolization of contaminated feces) were supposed to be the source of infection of severe acute respiratory syndrome (SARS) pandemic resulting in 329 infected individuals and 42 deaths (2003) (Wang et al. 2005a, b). By considering the view that SARS-CoV-2 remains viable in different water matrices (simulated and natural, Table S1 in supporting information), there is a chance that live infectious viruses can enter in WWTPs; nevertheless, their fate was decided by the used treatment processes and the survival efficacy of SARS-CoV-2. The detailed description of virus reduction/inactivation by different treatment processes used in WWTPs is shown in Table S3 (supplementary information). In addition, reduction in WWTPs was also influenced by the virus type (enveloped or non-enveloped) and its concentration (PFU and TCID50), disinfection ability, and inactivation rate. Thus, there is a probability that SARS-CoV-2 may enter in the river via effluent of WWTP and pose a human health risk upon accidental ingestion during swimming. The risk of human infection depends upon the (i) live infectious concentration of SARS-CoV-2 in the effluent of WWTP (removed during primary, secondary, and tertiary treatment), (ii) reduction of the virus by decay and dilution between the time of introduction at wastewater outfall and the time of encounter by a downstream recreational user (Haas 1983). The above discussion indicates that studies have been putting immense efforts on monitoring activities which are important as it provides occurrence information. However, very few publications discuss about the possible risk of infection from environment matrices (river, lakes, and ponds) which is an immediate public concern (Elsamadony et al. 2021; Liu et al. 2020; Bilal et al. 2020) (Fig. 1). Figure S1 (in supplementary information) highlights the current state-of-the-art which places less focus on estimating risk and more on understanding the overall big picture for making public health decision-making.
From the foregoing discussion, it is clearly evident that knowledge is scant regarding the transmission of COVID-19 via the fecal-oral route. So, there is a need for systematically studying the fecal-oral transmission via wastewater-based epidemiology. By keeping this in mind, literature-based occurrence data (concentration of SARS-CoV-2 (genomic copies/mL) in the raw wastewater and secondary treated effluent of WWTPs from different countries) was used to obtain an initial understanding of the risk of infection from the water environment (river). For calculating the probability of risk of infection, first-occurrence data related to the raw wastewater containing SARS-CoV-2 was collected. Thereafter, virus reduction in WWTP and dilution at the time of introduction of WWTP effluent into the downstream river was taken into account. Second, literature-based statistics of enteric viruses was used for the formulation of the ratio of infectious virus to genomic copies of SARS-CoV-2. Third, in the absence of SARS-CoV-2 DRM, it was assumed that SARS-CoV-2 follows the DRMs of SARS-CoV-1 and MERS-CoV surrogate coronavirus (epidemiologic resemblance of both coronaviruses in different environments) and/or influenza A virus (the most infectious virus accountable for respiratory illness) (Table 2). The emphasis of this research is to conduct quantitative microbial risk assessment and estimate the human health risk of infection due to exposure of SARS-CoV-2 from the river (swimming) via DRMs of respiratory and surrogate coronaviruses.
water (swimming). For this purpose, (i) genomic copies of SARS-CoV-2 present in the raw wastewater and secondary treated effluent of WWTP were gathered from the literature (details mentioned in “Exposure assessment”), (ii) concentration values of infectious SARS-CoV-2 from total genomic copies was computing using enteric virus information from cited articles (Table S3 in supporting information) for risk determination due to ingestion of SARS-CoV-2 (per dip/event), and (iii) different known DRMs of surrogate coronaviruses and influenza virus A (H5N1) were used in the absence of DRMs for SARS-CoV-2 (responsible for COVID-19 disease). The detailed justification for choosing the DRMs of surrogate coronaviruses and influenza virus A (H5N1) is presented. The QMRA steps used in the present work are shown below in a systematic manner.
Hazard identification
SARS-CoV-2 is a single-stranded RNA-based virus and belongs to a Betacoronavirus genus based on sequence identity (Marty and Jones 2020). SARS-CoV-2 is distantly related to human coronaviruses (229E, OC43, NL63, HKU1) which is accountable for 15–30% of cases of common cold and respiratory illness comprehensively (Mesel-Lemoine et al. 2012). The case fatality risk for COVID-19 has ranged between 3.4 and 8.4%, comparatively lower than SARS (up to 50%) and MERS (34–69%) (Jung et al. 2020; Park et al. 2018; Wang et al. 2020a, b, c). The infectivity potential of SARS-CoV-2 was estimated to be 1.4–6.5 cases. The virus is also responsible for GI tract infections (n = 103, diarrhea (34%), vomiting (4%), lack of appetite (79%), abdominal pain (2%)), impaired liver function (n = 95, 32.6%), and loss of smell (n = 103, 61%) (Lin et al. 2020; Ng and Tilg 2020; Yang and Tu 2020).
Exposure assessment
Possible scenarios from where humans get exposed to SARS-CoV-2 from environmental water are shown in Fig. 1. Nevertheless, this study focused on inadvertent ingestion of water during recreational activity (swimming) as an exposure scenario (ingestion = 32 mL per dip/swim event, Dufour et al. 2017) to provide a preliminary risk estimate (P(response) = risk of infection due to a single exposure event) based on limited information. Exposure through other potential routes (such as dermal and inhalation) was not considered here.
For exposure assessment, the concentrations of SARS-CoV-2 in raw wastewater and secondary treated effluent of WWTP were gathered from the literature (Table 1) and used for QMRA analysis. SARS-CoV-2 concentration from secondary treated effluent of WWTP was considered here to check how much risk was there if a considerate value of the virus was present after the secondary treatment. Nevertheless, the reported values in the literature tell us the measurable viral load above the detection limit. But this picture is not appropriate to describe the viral load in the effluent, and the observations under the detection limit should be taken into account to avoid overestimation of the risk. In order to avoid this situation, the detection limit of the instrument (RT-PCR in most of the cases) cited in the research articles was substituted for the below detection values. The detailed information about the river water considered here is presented in estimation of virus concentration in wastewater and river water. The frequency of exposure was considered to be a single event, since the emphasis of this study is to calculate risks at the recreational sites (river) during the COVID-19 outbreak rather than extrapolating annual risks.
Calculation of ratio of infectious virus concentration to total virus particle concentration
The ratio of infectious to total virus particles with regard to SARS-CoV-2 was not available at the time of submission of this paper. Outcomes of previous studies were used for formulating the ratio of infectious to the total number of virus particles for calculating the theoretical concentration values of SARS-CoV-2 and are mentioned below (Table 2).
The relationship between genomic copies (total RNA virus) and PFU or TCID50 (infectious live virus) of SARS-CoV-2 is not available; hence, enteric virus information from the literature was used for the calculation of concentration values of infectious (PFU) versus total genomic copies (gc) of SARS-CoV-2. The relationship of this ratio for SARS-CoV-2 and surrogate virus depends on the (i) interrelation between SARS-CoV-2 and surrogate coronavirus and (ii) concentration of infectious virus out of the total genomic copies of surrogate coronavirus. More work is needed to experimentally determine this information, and then, the ratio of the concentration of infectious virus to the total screened genomic copies of SARS-CoV-2 can be accurately linked with that of the surrogate viruses.
Estimation of virus concentration in wastewater and river water
The risk of human infection depends upon the (i) live infectious concentration of SARS-CoV-2 in the effluent of WWTP, (ii) reduction of the virus by decay and dilution between the time of introduction at wastewater outfall and the time of encounter downstream in a river (Haas 1983). The concentration (Nie) of SARS-CoV-2 in the effluent of WWTP was calculated by Eq. (1):
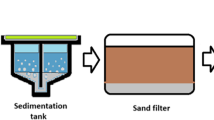
where Nio denotes the concentration of ith (SARS-CoV-2 in this case) type of virus in the raw wastewater, fi2 and fid are the fractions of microorganisms (viruses are considered here: Adenoviruses, Enteroviruses, Rotavirus, Noraviruses, and Astroviruses) removed during (i) primary and secondary treatment and (ii) disinfection (in the absence of disinfection, fid = 0). Here, in the present investigation, the activated sludge process was assumed as a secondary treatment method in the WWTP and chlorination as a tertiary treatment. The considered values for fi2 and fid were 0.97 (97% removal efficiency) and 0.99 (99% removal efficiency), respectively. The percentage values of removal efficiencies were collected from the literature and their median values were used in the calculation of Nie (Table S3 in supporting information) (Le Cann et al. 2004; Kuo et al. 2010; Francy et al. 2012; Zhou et al. 2015). Also, a complex series of reactions take place in the WWTP under a natural ambient environment such as endogenous decay of virus, sedimentation and scouring, light-mediated inactivation, reproduction, and predation. The downstream concentration of the SARS-CoV-2 virus (Nit) at a travel time “t” from the discharge was calculated using Eq. (2):
where ki is the decay coefficient of the virus within the reach of interest, t is the travel time, and D is the dilution factor (ratio of upstream to effluent volumetric flow)
In the present study, ki = 0.69/day, t = 2 days and D = 99:1 were used for risk calculation henceforth (adapted from Haas 1983). While calculating Nit, three suppositions were used: (i) uniform, plug flow conditions at steady state in the river under consideration, (ii) decay constant value was fixed in the river, and (iii) the dilution of the WWTP effluent into the stream occurred once instantly at the outfall, and thereafter, no further dilution takes place in the river. The model estimate of concentration of SARS-CoV-2 in river water (Nit) was used for estimating the probability of infection (shown in the subsequent section) due to inadvertent ingestion of river water by recreators.
Dose-response model
Due to the unavailability of explicit DRM for SARS-CoV-2, this study assumes that DRM of SARS-CoV-2 is the same as D-R models of SARS with different characteristics (Table 3) (Haas 2020). This study hypothesized that SARS-CoV-2 DRM is similar to the (i) SARS-CoV-1 (DRM-1), (ii) SARS-CoV-1 (DRM-2), (iii) MERS-CoV (DRM-3), and (iv) influenza A (H5N1) (DRM-4). These models were selected based on the similar epidemiological character in different environments (best available coronavirus surrogates) and pathogenicity. In contrast, another supposition is that the surrogate coronavirus DRMs can be used for the SARS-CoV-2 after using a proposed conversion parameter factor (f) and included in the expression for estimating risk.
Estimation of risk of infection
The risk of infection was calculated using each of these four DRMs (i.e., four scenarios). Each of these models has an exponential form as shown below:
where P = probability of infection after a single exposure at the dose “d”, i.e., ingestion dose per event (ingestion volume = 32 mL in the present investigation) which were capable of initiating infection, k is the probability that one particulate will initiate infection and the value of “k” depends upon the DRM selected (mentioned in Table 3). Here, f was assumed to be 1 as currently experimental data is not available in the literature to determine the value of this parameter. More work is needed in this regard. Upon getting the f value from experimental studies, the risk estimate can be updated.
One-way sensitivity analysis was done to understand the effect of variation of one parameter (from low to high value) on the output variable, i.e., risk of infection (keeping other variables at their mean values). This exercise helped in identifying parameters which might be contributing more to the uncertainty in risk of infection values. The following parameters were varied one at a time while other parameters were kept fixed at their mean values for estimating the risk of infection: (i) fraction of viruses removed after activated sludge process and disinfection unit (chlorination), (ii) fraction of infectious concentration of SARS-CoV-2 present in the total screened genomic copies, (iii) “k” value (DRM), and (iv) molecular concentration of SARS-CoV-2 (gc/mL)). The sensitivity index (SI) for a given parameter was calculated using Eq. (4) where information related to values of risk of infection at high and low values of parameters and low, high, and median values of the parameter under consideration was used. In Eq. (4), PhighX and PlowX indicate risk of infection values calculated at high value of parameter ChighX) and low value of parameters (ClowX). Also, CaveX indicates the average value of the considered parameter. All variables were ordered in decreasing order of their SI values for identifying parameters which contribute more to the uncertainty in risk of infection value.
Results and discussion
Roadmaps mentioned in Fig. 2 explain the steps used in the calculation of the theoretical concentration of SARS-CoV-2 in river water after effluent being discharged from the WWTPs and human risk of infection owing to the presence of SARS-CoV-2 in environmental water.
Risk estimation in river water at the point of discharge from WWTP
Case 1: Risk estimation using raw wastewater containing SARS-CoV-2 after secondary treatment and disinfection in wastewater plant and dilution in river
Table 4 shows the literature-reported concentration (genomic copies/mL) of SARS-Co-V-2 in the raw wastewater (lowest value = 1.9 × 10−2 and highest value = 3 × 103). The calculated concentration values of SARS-CoV-2 (genomic copies/mL) in the mixed river water (after dilution, decay, and travel) at the point where WWTP effluent entered into the river stream were found to be 1.43 × 10−8 genomic copies/mL and 2.26 × 10−3 genomic copies/mL (corresponding to lowest and highest literature-reported values). The calculated ingested concentration values of SARS-CoV-2 after per dip were found to be 4.6 × 10−7 (lowest) and 7.25 × 10−2 genomic copies/event (highest) (Table 4, column E) and further used for the calculation of the infectious concentration of SARS-CoV-2.
The calculated infectious concentration values of SARS-CoV-2 was used for risk estimation and risk values lie in between 1.7 × 10−8 and 9.0 × 10−12 per event (lowest reported value) and 7.9 × 10−4 to 8.0 × 10−6 per event (highest reported value) for all the considered DRMs (Table 5). However, for DRM-4, the risk values were found to be on the higher side as compared to other DRMs.
Case 2: Risk estimate using secondary treated effluent (without disinfection) containing SARS-CoV-2 after dilution in river
The calculated concentration value of ingested virus after each dip was found to be 80.5 genomic copies/event (Table 4, column E). Risk of infection was noted at calculated infectious concentration values using considered DRMs and was observed to range from 8.8 × 10−4 to 5.8 × 10−1 per event (Table 5). The overall trend (based on high- to low-risk values) of the effect of DRMs on risk estimate was observed to be DRM-4 > DRM-3 > DRM-2 > DRM-1. Furthermore, additional information is mentioned in Table S4 (see supplementary information) where risk calculation was done by considering two cases: (i) when non-detects were replaced with the limit of detection and (ii) zero genomic copies/event. It was noticed that calculated risk values were almost equivalent to the allowable value of risk (1/10,000) (Table S4, see supplementary information) using four assumed D-R scenarios (Table 3).
The assumptions used in the present work were considered applicably conventional and protective to human health and supported by literature information. However, information is lacking regarding the SARS-CoV-2 in this regard. Hence, we considered enteric viruses responsible for GI tract illness and transmitted via the fecal-oral route and/or fomites for assumptions formulation (calculation of infectious virus concentration out of total genomic copies) (He and Jiang 2005; van Heerden et al. 2005a, b; de Roda Husman et al. 2009; Pinto et al. 2009; Rigotto et al. 2010; Aslan et al. 2011; McBride et al. 2013; Carducci et al. 2016). The noted risk values in the present work are sometimes 10,000 (Table 5) times higher than the USEPA standard risk value for pathogens (1/10,000) emphasizing that extra safeguards are prerequisite during COVID-19 outbreaks for recreational activities. Scientists across the globe work comprehensively regarding the occurrence (Table 1), molecular detection (viral RNA, RT-PCR, qPCR; Table 1), and medicine formulation for SARS-CoV-2; nevertheless, still many citations were not able regarding the human health risk due to consumption of SARS-CoV-2 via drinking and swimming and through contaminated surfaces and other occupational works (La Rosa et al. 2020; Nemudryi et al. 2020; Randazzo et al. 2020; Rimoldi et al. 2020; Wu et al. 2020; Wurtzer et al. 2020). Zaneti et al. (2020) published the first articles that studied the health risks of SARS-CoV-2 present in the raw wastewater to WWTP workers and risk values are 1.03 × 102 to 1.31 × 104 genomic copies/mL (0.1 to 13.06 PFU/mL, respectively) (Table 6). Further, Shutler et al. (2020) identified the risk of infection due to SARS-CoV-2 present in the polluted water systems. They suggested that the level of risk (high, medium, and low) depends upon the river dilution, geographic location, temperature, and water usage capacity. They used literature on the number of infectious Adenovirus copies in sewage and wastewater discharge into rivers to select high (10−1), medium (10−2), and low (10−3) estimates for the ratio of infectious virus to genome copies (Table 6). However, they have not used these observations further for risk estimation.
Effect of “k” value on risk estimate
The DRMs used were based on the exponential model. As per the Haas et al. (1999) study for the usage of exponential models, microorganisms (SARS-CoV-2 in the present case) are randomly distributed in the water and follow Poisson distribution for infection to occur and the probability of infection per ingested or inhaled organism is constant. Each microorganism has the same fixed probability (“r” or “k”) of surviving and reaching a host site at which infection may result. So, based on this assumption, risk estimation was largely dependent on the value of “r” or “k.” Also, the results presented in Table 5 show a positive correlation with this statement because the number of viruses ingested was the same for all the studied DRMs. Figures S2, S3, S4, S5, and S6 in the supporting information show the influence of different “k” values on the risk of infection at three different virus concentrations (0.019 genomic copies/mL, 1000 genomic copies/mL, and 3000 genomic copies/mL). The noted risk values were significantly different and lie in between 9.0 × 10−12 and 5.8 × 10−1 per event.
Sensitivity analysis
The order of parameter affecting uncertainty in risk of infection value based on the sensitivity index (Table 7) was found to be (high to low SI value): Molecular concentration of SARS-CoV-2 (gc/mL) (rank 1) > “k” value (DRM) (rank 2) > fraction of infectious concentration of SARS-CoV-2 present in the total screened genomic copies (rank 3) > fraction of virus removal in activated sludge process ≈ fraction of virus removal in disinfection unit (chlorination). It indicates that more information is required for properly characterizing uncertainty in these parameters so that uncertainty in the value of risk of infection can be properly characterized.
Summary and conclusions
The present study investigated the risk of infection due to the accidental ingestion of river water during swimming (32 mL per dip/event) which hypothetically receives SARS-CoV-2 virus-containing effluent of WWTP. The summary of the noted results are:
-
1.
The calculated concentration values of ingested SARS-CoV-2 ranged between 4.6 × 10−7 and 80.5 genomic copies/dip or /event.
-
2.
Risk values (> 9 × 10−12 to 5.8 × 10−1) was found to be > 1/10,000 risk of infection, a commonly cited benchmark of concern for microbial risk due to water (Regli et al. 1991), for all the studied DRM scenarios, indicating a chance of health risks.
-
3.
It was noted that the risk of infection was directly proportional to the “k” value of DRMs irrespective of the concentration of ingested virus.
-
4.
The overall trend of uncertainties was (in order of importance): Molecular concentration of SARS-CoV-2 (gc/mL) > “k” value (DRM) > fraction of infectious concentration of SARS-CoV-2 present in the total screened genomic copies > fraction of virus removal in activated sludge process ≈ fraction of virus removal in disinfection unit (chlorination).
In a nutshell, information regarding infectious concentration (PFU and TCID50) of SARS-CoV-2 is missing (because of the unavailability of surrogate viruses’ information) and requires a more intensive emphasis in the forthcoming studies, so that in the future, there will more data and DRM to calculate the accurate human risk of infection. This work will give insights for risk quantification till the DRM for SARS-CoV-2 is available. Additionally, it helps legislators and other regulatory bodies (WHO and USEPA) for taking appropriate measures before discharging the WWTPs’ effluent in the environmental water especially the ones used for recreational activities. However, the current finding suggests that fecal-oral transmission is not proven; an analogy with the previous study related to MERS and SARS suggests that it could be a potential source of risk if studied in detail.
Overall, there is a need for initiating concerted efforts for (i) method development for determining the infectious virus concentration using a molecular-based method like integrated cell culture PCR (PFU or TCID50 versus genomic copies) (Fig. 3, challenges and key perspective (CKP) 1), (ii) pursuing long-term continuous monitoring of infectious virus concentrations in community-based wastewater effluent (Fig. 3, CKP 1 and 2). Also, focus is required for standardizing the dose-response relationship using known concentrations of doses (PFU) and for determining its susceptibility by animal models as well as epidemiological studies.
The diagram illustrates the inter-linkages of different factors for elucidating human health risks. The light green color dashed box shows the challenges and key perspective which needs attention. WWTP, wastewater treatment plant; CHP, challenges and key perspective; DRM, dose-response model; PFU, plaque-forming unit; TCID50, median tissue culture infectious dose
Data availability
All the data involved in this manuscript can be obtained online (supporting information). No additional data available.
References
Ahmed W, Angel N, Edson J, Bibby K, Bivins A, Brien JWO, Choi PM, Kitajima M, Simpson SL, Li J, Tscharke B, Verhagen R, Smith WJM, Zaugg J, Dierens L, Hugenholtz P, Thomas KV, Mueller JF (2020) First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Science of the Total Environment. 728:138764. https://doi.org/10.1016/j.scitotenv.2020.138764
Albuquerque ND, Baig E, Ma X, Zhang J, He W, Rowe A, Habal M, Liu M, Shalev I, Downey GP, Gorczynski R, Butany J, Leibowitz J, Weiss SR, McGilvray ID, Phillips M, Fish MJ, Levy GA (2006) Murine hepatitis virus strain 1 produces a clinically relevant model of severe acute respiratory syndrome in a/j mice. Journal of Virology. 80:10382–10394
Aslan A, Xagoraraki I, Simmons FJ, Rose JB, Dorevitch S (2011) Occurrence of Adenovirus and other enteric viruses in limited-contact freshwater recreational areas and bathing waters. Journal of Applied Microbiology. 111:1250–1261. https://doi.org/10.1111/j.1365-2672.2011.05130.x
Bilal M, Nazir MS, Rasheed T, Parra-Saldivar R, Iqbal HMN (2020) Water matrices as potential source of SARS-CoV-2 transmission–an overview from environmental perspective. Case Studies in Chemical and Environmental Engineering 2:100023
Carducci A, Donzelli G, Cioni L, Verani M (2016) Quantitative microbial risk assessment in occupational settings applied to the airborne human adenovirus infection. International Journal of Environmental Research and Public Health. 13:733–743. https://doi.org/10.3390/ijerph13070733
Chen N, Zhou M, Dong X, Qu J, Gong F, Han Y, Qiu Y, Wang J, Liu Y, Wei Y, Xia J, Yu T, Zhang X, Zhang L (2020) Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet. 395(10223):507–513. https://doi.org/10.1016/S0140-6736(20)30211-7
de Roda Husman AM, Lodder WJ, Rutjes SA, Schijven JF, Teunis PFM (2009) Long-term inactivation study of three enteroviruses in artificial surface and ground waters, using PCR and cell culture. Applied and Environmental Microbiology. 75:1050–1057
DeDiego ML, Pewe L, Alvarez E, Rejas MT, Perlman S, Enjuanes L (2008) Pathogenicity of severe acute respiratory coronavirus deletion mutants in hACE-2 transgenic mice. Virology. 376:379–389
Dufour P, Behymer TD, Cantu R, Magnuson M, Wymer LJ (2017) Ingestion of swimming pool water by recreational swimmers. Journal of Water and Health. 15:429–437
Elsamadony M, Fujii M, Miura T, Watanabe T (2021) Possible transmission of viruses from contaminated human feces and sewage: implications for SARS-CoV-2. Science of the Total Environment 755:142575
Francy DS, Stelzer EA, Bushon RN, Brady AMG, Williston AG, Riddell KR, Borchardt MA, Spencer SK, Gellner TM (2012) Comparative effectiveness of membrane bioreactors, conventional secondary treatment, and chlorine and UV disinfection to remove microorganisms from municipal wastewaters. Water Research. 46:4164–4178
Guan W, Ni Z, Hu Y, Liang W, Ou C, He J, Liu L, Shan H, Lei C, Hui D, Du B, Li L, Zeng G, Yuen K, Chen R, Tang C, Wang T, Chen P, Xiang J et al (2020) Clinical characteristics of coronavirus disease 2019 in China. The New England Journal of Medicine. 382:1708–1720. https://doi.org/10.1056/NEJMoa2002032
Haas CN (1983) Effect of effluent disinfection on risks of viral disease transmission via recreational water exposure. Journal of the Water Pollution Control Federation. 55(8):1111–1116
Haas CN (2020) Coronavirus and environmental engineering science. Environmental Engineering and Science. 37:1–2
Haas CN, Rose JB, Gerba CP (1999) Quantitative microbial risk assessment. John Wiley and Sons, New York
Haramoto E, Malla B, Thakali O, Kitajima M (2020) First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Science of the Total Environment. 737:140405. https://doi.org/10.1016/j.scitotenv.2020.140405
He J-W, Jiang S (2005) Quantification of Enterococci and human Adenoviruses in environmental samples by real-time PCR. Applied and Environmental Microbiology. 71(5):2250–2255
Heller L, Mota CR, Greco DB (2020) COVID-19 faecal-oral transmission: are we asking the right questions? Science of the Total Environment. 729:1–3. https://doi.org/10.1016/j.scitotenv.2020.138919
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, Zhang L, Fan G, Xu J, Gu X, Cheng Z, Yu T, Xia J, Wei Y, Wu W, Xie X, Yin W, Li H, Liu M et al (2020) Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 395:497–506. https://doi.org/10.1016/S0140-6736(20)30183-5
Jung S-M, Akhmetzhanov AR, Hayashi K, Linton NM, Yang Y, Yuan B, Kobayashi T, Kinoshita R, Nishiura H (2020) Real-time estimation of the risk of death from novel coronavirus (COVID-19) infection: inference using exported cases. Journal of Clinical Medicine. 9:523. https://doi.org/10.3390/jcm9020523
Kitajima M, Ahmed W, Bibby K, Carducci A, Gerba CP, Hamilton KA, Haramoto E, Rose JB (2020) SARS-CoV-2 in wastewater: state of the knowledge and research needs. Science of the Total Environment. 739:139076. https://doi.org/10.1016/j.scitotenv.2020.139076
Kocamemi BA, Kurt H, Hacıoglu S, Yarali C, Saatci AM, Pakdemirli B (2020) First data-set on SARS-CoV-2 detection for Istanbul wastewaters in Turkey. MedRxiv2020.05.03.20089417. https://doi.org/10.1101/2020.05.03.20089417
Kuo DHW, Simmons FJ, Blair S, Hart E, Rose JB, Xagoraraki I (2010) Assessment of human adenovirus removal in a full-scale membrane bioreactor treating municipal wastewater. Water Research. 44:1520–1530
La Rosa G, Bonadonna L, Lucentini L, Sebastien K, Suffredini E (2020) Coronavirus in water environmental: Occurrence, persistence and concentration methods-a scoping review. Water Research. 179:115899. https://doi.org/10.1016/j.watres.2020.115899
Le Cann P, Ranarijaona S, Monpohe S, Monpoeho S, Le G, Ferre V (2004) Quantification of human astroviruses in sewage using real time RT-PCR. Microbiological Research. 155:11–15
Lin L, Jiang X, Zhang Z, Huang S, Zhang Z, Fang Z, Gu Z, Gao L, Shi H, Mai L, Liu Y, Lin X, Lai R, Yan Z, Li X, Shan H (2020) Gastrointestinal symptoms of 95 cases with SARS-CoV-2 infection. Gut. 69:997–1001
Liu D, Thompson JR, Carducci A, Bi X (2020) Potential secondary transmission of SARS-CoV-2 via wastewater. Science of the Total Environment. 749:142358
Lunn TJ, Restif O, Peel AJ, Munster VJ, De Wit E, Sokolow S, Van Doremalen N, Hudson P, McCallum H (2019) Dose-response and transmission: the nexus between reservoir hosts, environment and recipient hosts. Philosophical Transactions of the Royal Society B: Biological Sciences 374:20190016. https://doi.org/10.1098/rstb.2019.0016
Marty AM, Jones MK (2020) The novel coronavirus (SARS-CoV-2) is a one health issue. One Health. 9:100123
McBride GB, Stott R, Miller W, Bambic D, Wuertz S (2013) Discharge-based QMRA for estimation of public health risks from exposure to stormwater-borne pathogens in recreational waters in the United States. Water Research. 47:5282–5297. https://doi.org/10.1016/j.watres.2013.06.001
Medema, G., Heijnen, L., Elsinga, G., Italiaander, R. and Brouwer, A. (2020). Presence of SARS-Coronavirus-2 in sewage. medRxiv. 2020.03.29.20045880. https://doi.org/10.1101/2020.03.29.20045880.
Mesel-Lemoine M, Millet J, Vidalain P-O, Law H, Vabret A, Lorin V, Escriou N, Albert ML, Nal B, Tangy F (2012) A human coronavirus responsible for the common cold massively kills dendritic cells but not monocytes. Journal of Virology. 86:7577–7587. https://doi.org/10.1128/jvi.00269-12
Nemudryi A, Nemudraia A, Surya K, Wiegand T, Buyukyoruk M, Wilkinson R, Wiedenheft B (2020) Temporal detection and phylogenetic assessment of SARS-CoV-2 in municipal wastewater. medRxiv. https://doi.org/10.1101/2020.04.15.20066746
Ng SC, Tilg H (2020) COVID-19 and the gastrointestinal tract: more than meets the eye. Gut. 69:973–974
Park JE, Jung S, Kim A (2018) MERS transmission and risk factors: a systematic review. BMC Public Health. 18(1):574. https://doi.org/10.1186/s12889-018-5484-8
Pinto RM, Costafreda MI, Bosch A (2009) Risk assessment in shellfish-borne outbreaks of hepatitis A. Applied and Environmental Microbiology. 75:7350–7373
Randazzo, W., Truchado, P., Ferrando, E.C., Simon, P., Allende, A. and Sanchez, G. (2020). SARS-CoV-2 RNA titers in wastewater anticipated COVID-19 occurrence in a low prevalence area. medRxiv. 2020.04.22.20075200. https://doi.org/10.1101/2020.04.22.20075200.
Regli S, Rose JB, Haas CN, Gerba CP (1991) Modeling the risk from Giardia and viruses in drinking water. Journal of American Water Works Association. 83:76–84
Rigotto CM, Victoria M, Moresco V, Kolesnikovas CK, Correa AA, Souza DSM, Miagostovich MP, Simoes CMO, Barardi CRM (2010) Assessment of adenovirus, hepatitis A virus and rotavirus presence in environmental samples in Florianopolis. South Brazil. Journal of Applied Microbiology. 109:1979–1987
Rimoldi SG, Stefani F, Gigantiello A, Polesello S, Comandatore F, Mileto D, Maresca M, Longobardi C, Mancon A, Romeri F, Pagani C, Moja L, Gismondo MR, Salerno F (2020) Presence and vitality of SARS-Cov-2 virus in wastewaters and rivers. medRxiv. https://doi.org/10.1101/2020.05.01.20086009
Sherchan SP, Shahin S, Ward LM, Tandukar S, Aw TG, Schmitz B, Ahmed W, Kitajima M (2020) First detection of SARS-CoV-2 RNA in wastewater in North America: a study in Louisiana, USA. Science of the Total Environment. 743:140621. https://doi.org/10.1016/j.scitotenv.2020.140621
Shutler J, Zaraska K, Holding T, Machnik M, Uppuluri K, Ashton I, Migdal L, Dahiya R (2020) Risk of SARS-CoV-2 infection from contaminated water systems. medRxiv. https://doi.org/10.1101/2020.06.17.20133504
van Heerden J, Ehlers MM, Grabow WOK (2005a) Detection and risk assessment of adenoviruses in swimming pool water. Journal of Applied Microbiology. 99:1256–1264
van Heerden J, Ehlers MM, Heim A, Grabow WOK (2005b) Prevalence, quantification and typing of adenoviruses detected in river and treated drinking water in South Africa. Journal of Applied Microbiology. 99:234–242
Wang XW, Li JS, Guo TK, Zhen B, Kong QX, Yi B, Li Z, Song N, Jin M, Xiao WJ, Zhu XM, Gu CQ, Yin J, Wei W, Yao W, Liu C, Li JF, Ou GR, Wang MN et al (2005a) Concentration and detection of SARS coronavirus in sewage from Xiao Tang Shan Hospital and the 309th hospital. Journal of Virological Methods. 128:156–161. https://doi.org/10.1016/j.jviromet.2005.03.022
Wang XW, Li JS, Jin M, Zhen B, Kong QX, Song N, Xiao WJ, Yin J, Wei W, Wang GJ, Si BY, Guo BZ, Liu C, Ou GR, Wang MN, Fang TY, Chao FH, Li JW (2005b) Study on the resistance of severe acute respiratory syndrome-associated coronavirus. Journal of Virological Methods. 126:171–177. https://doi.org/10.1016/j.jviromet.2005.02.005
Wang C, Horby PW, Hayden FG, Gao GF (2020a) A novel coronavirus outbreak of global health concern. Lancet. 395:470–473. https://doi.org/10.1016/S0140-6736(20)30185-9
Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z (2020b) Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. Journal of American Medical Association. 323:1061–1069. https://doi.org/10.1001/jama.2020.1585
Wang W, Xu Y, Gao R, Lu R, Han K, Wu G, Tan W (2020c) Detection of SARS-CoV-2 in different types of clinical specimens. Journal of American Medical Association. 323:1843–1844. https://doi.org/10.1001/jama.2020.3786
Watanabe T, Bartrand TA, Weir MH, Omura T, Haas CN (2010) Development of a dose-response model for SARS coronavirus. Risk Analysis. 30:1129–1138. https://doi.org/10.1111/j.1539-6924.2010.01427.x
World Health Organization. (2020a). Pneumonia of unknown cause-China [WWW document]. URL. https:// www.who.int/csr/don/05-january-2020-pneumonia-of-unkown-cause-china/en/.
World Health Organization. (2020b). Statement on the second meeting of the international health regulations (2005) emergency committee regarding the outbreak of novel Coronavirus (2019-nCoV).
Wu F, Xiao A, Zhang J, Gu X, Lee W, Kauffman K, Hanage W, Matus M, Ghaeli N, Endo N, Duvallet C, Moniz K, Erickson T, Chai P, Thompson J, Alm E (2020) SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases. medRxiv. https://doi.org/10.1101/2020.04.05.20051540
Wurtzer S, Marechal V, Mouchel J, Moulin L (2020) Time course quantitative detection of SARS-CoV-2 in Parisian wastewaters correlates with COVID-19 confirmed cases. medRxiv. https://doi.org/10.1101/2020.04.12.20062679
Xiao F, Tang M, Zheng X, Li C, He J (2020) Evidence for gastrointestinal infection of SARS-CoV-2. medRxiv. https://doi.org/10.1101/2020.02.17.20023721
Yang L, Tu L (2020) Implications of gastrointestinal manifestations of COVID-19. The Lancet Gastroenterology and Hepatology. 5:629–630. https://doi.org/10.1016/S2468-1253(20)30132-1
Zaneti RN, Girardi V, Spilki FR, Mena K, Westphalen APC, Colares ER d C, Pozzebon AG, Etchepared RG (2020) QMRA of SARS-CoV-2 for workers in wastewater treatment plants. medRxiv. https://doi.org/10.1101/2020.05.28.20116277
Zhou J, Wang XC, Ji Z, Xu LM, Yu ZZ (2015) Source identification of bacterial and viral pathogens and their survival/fading in the process of wastewater treatment, reclamation, and environmental reuse. World Journal of Microbial Biotechnology 31:109–120 Weblink:Qmrawiki: http://qmrawiki.org/experiments/escherichia-coli. Accessed on 26-07-2020
Acknowledgements
The authors would also like to acknowledge Drexel University, USA, and the Department of Civil Engineering, IIT Delhi, for supporting this study.
Author information
Authors and Affiliations
Contributions
Neha Tyagi: Conceptualization, methodology, result analysis, data curation, writing—original draft
Patrick L Gurian: Conceptualization, review and editing, visualization
Arun Kumar: Conceptualization, writing—review and editing, visualization, supervision
All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable here.
Consent for publication
The authors declare that there is no conflict of interest regarding the publication of this paper.
Competing interests
The authors declare no competing interests
Additional information
Responsible Editor: Lotfi Aleya
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• Study estimated the human health risk due to ingestion of SARS-CoV-2 via swimming
• Literature-based enteric virus information was used for ratio formulation of infectious virus to genomic copies
• Concentration of ingested SARS-CoV-2 ranged from 4.6 × 10−7 to 80.5 genomic copies/swim
• Risk values (per event) lie in between 9.0 × 10−12 and 5.8 × 10−1
• Risk estimation was largely dependent on the molecular concentration of SARS-CoV-2
Supplementary information
ESM 1
(DOCX 219 kb)
Rights and permissions
About this article
Cite this article
Tyagi, N., Gurian, P.L. & Kumar, A. Using QMRA to understand possible exposure risks of SARS-CoV-2 from the water environment. Environ Sci Pollut Res 29, 7240–7253 (2022). https://doi.org/10.1007/s11356-021-16188-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-16188-0